I had the pleasure this June of presenting my class’s bioinformatics project to attendees of this year’s SEA-PHAGES symposium. Our project was rather abstract, relying on frequency analysis of DNA n-tuples to infer relationships between genomes, and many of the symposium attendees had minimal experience with the specifics of what we were presenting. It was a unique opportunity to see if a visual explanation would help convey what’s a rather technical and non-intuitive topic.
Elements
I took advantage of my medium—relying heavily on animation and colour to convey points that I would otherwise have to resort to jargon to explain—


I also used animation to convey context and relevance—particularly important in a math-heavy project:

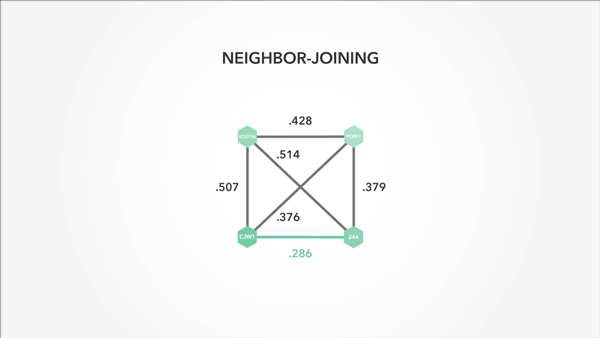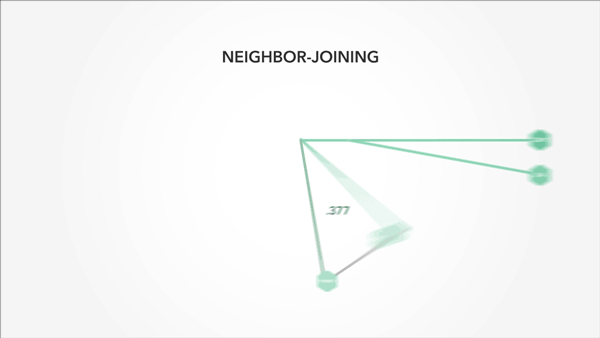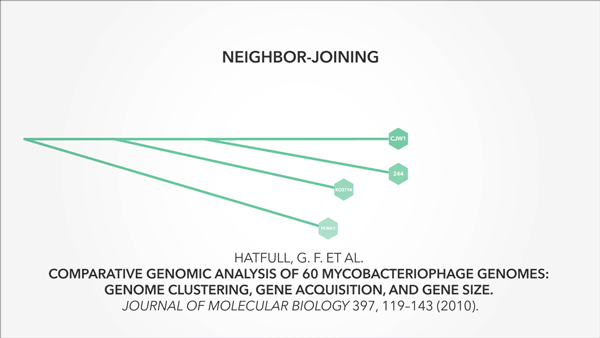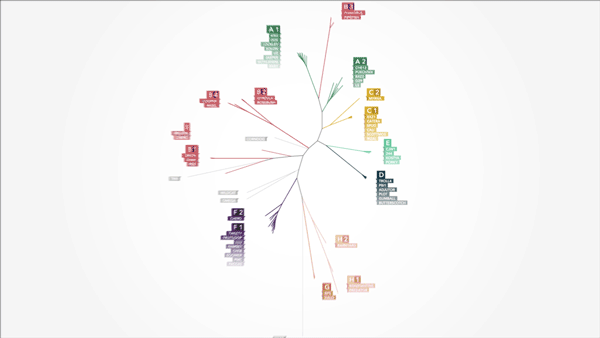
Animation that is precise and understated is startlingly effective at getting ideas across.
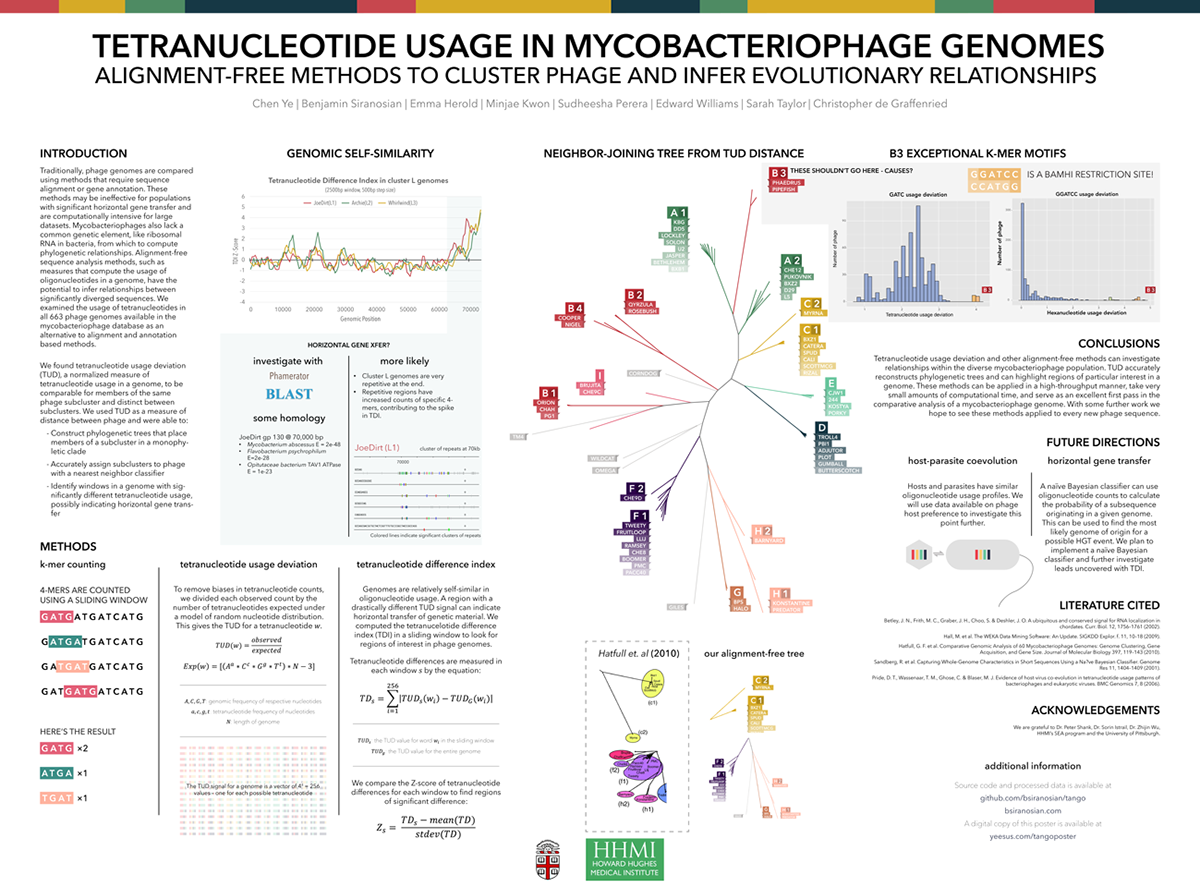

Color was equally important, allowing me to clearly group and organize distinct objects. This was very important for our phylogenetic tree, where cluster relationships between the 60 bacteriophages could clearly be determined based on color.
Using color and motion to convey points allows for more direct communication of concepts—we can process visual information much faster and more intuitively than we can parse text.
Process
On the technical side, my workflow can be broken down into four main components—narrative building, scene construction (in Illustrator), animation (in After Effects), packaging (in PowerPoint). A brief summary of each follows:
the narrative:
Arguably the most important step in the process—I’ll draft out a rough script, starting with an outline, and eventually refining that into prose addressing each point I would like to convey. I tend not to use visual storyboards, so getting the narrative right is essential.
a slide in Adobe Illustrator. note that I needed to break up animated elements into different layers
Next, I’ll break up the narratives into a series of slides I would like to illustrate. This is a highly subjective task, and slides can be highly simple (static text on a screen), or incredibly complex (animated tree construction). Whatever it is, it gets drawn out in Illustrator, and broken up into layers in preparation for animation.
the final set of slides I created for tango
animation:
Here’s the hard and time-consuming part—animating it all in After Effects. My workflow has been significantly sped up in the past year due to my increased use of expressions and scripts, but due to the multi-purpose nature of After Effects, much of the animation work is still rather manual and low-level. Animation generally comprises a slide transition—usually panning or zooming, and an internal animation sequence. The average slide takes about an hour to create and completely animate.
powerpoint; packaging:
After all this, I’ve got a composition in After Effects that I can render out to a video. I bring this video into PowerPoint, and use PowerPoint’s bookmark feature to bookmark spots in the video I would like the presentation to pause at.
PowerPoint’s video bookmark feature. 1 bookmark per slide pause is needed.
From there, it’s a simple (if tedious) step to add a pause trigger to every bookmark. Once that (and adding play triggers to the next action) is complete, the PowerPoint will behave like a normal (albeit incredibly large) presentation, advancing slide whenever ‘next’ is hit.
This is a simplified overview of my workflow in creating these kinds of presentations.
You can look at the raw files themselves in our Github repository, as well as read more about the science behind things at Ben’s blog.






